 Explore our latest work
Explore our latest work

This blog series explores how biological neural networks can be integrated into modern AI systems. The posts progress from practical applications to increasingly general principles, moving from perception to representation, dynamics, and ultimately algorithm discovery. This post focuses on how AI models process visual information, showing how biologically derived neural connectivity patterns can be harnessed in AI models to improve image reconstruction.
Note: This is a living document, and we are building in the open. We expect frequent updates and actively welcome feedback from the community.
For the first time, we show that adding a small, biologically inspired software component to a modern AI image model improves image detail and quality without retraining the entire system. This result translates directly to better performance in products such as image compression, interactive games, simulations, robotics training, virtual worlds, and world models. Importantly, our results also suggest that neural-based components can reduce the computing power needed to generate high-quality images, offering a practical way to learn from how real neural systems organize and process information and to use those insights to discover new ways for AI models to learn.
Adding a small, biologically inspired software add-on to an existing AI image compression model produces noticeably sharper and more detailed image reconstructions. The improvement comes without retraining the full model and with only a minimal increase in complexity, making it practical for real-world use.

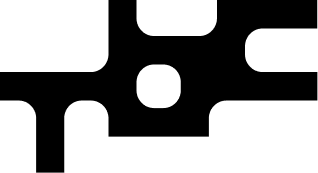
This figure compares an original image with two reconstructed versions. The middle image shows the result from a modern AI image compression model, which loses fine detail. The image on the right shows the same model with a small, biologically inspired software add-on, which preserves sharper structure and visual clarity. The biological add-on improves image quality while adding a negligible amount of additional model complexity.
We invite teams interested in new ways of building AI systems to connect with us. We are exploring how biologically inspired software can improve learning efficiency, reduce compute cost, and unlock new capabilities in modern AI. If you are thinking about what the next generation of computing systems could look like, we would love to explore this together.
Modern visual AI systems face a persistent tradeoff between image quality and computational cost. High-fidelity image generation and reconstruction are possible when models are allowed to compute extensively, but these approaches quickly become impractical for real-time or interactive settings. Systems designed to operate efficiently under tight latency and compute constraints often sacrifice fine detail and visual realism, limiting their usefulness in applications such as games, simulations, robotics, and virtual environments.
Variational autoencoders (VAEs) sit at the center of this challenge. By compressing high-dimensional images into compact latent representations, VAEs enable scalable visual processing but introduce a bottleneck that can degrade reconstruction quality. Common failure modes include over-smoothing, loss of fine structure, and sensitivity to distribution shift, particularly when models are pushed beyond the operating regimes they were optimized for. Improving reconstruction fidelity typically requires increasing model capacity or retraining, both of which raise compute costs and reduce deployability.
Biological visual systems offer a useful reference point for addressing this challenge. Mammalian neural circuits process complex visual scenes in real time while operating under strict energy constraints, relying on local connectivity and distributed encoding to preserve structure and robustness. In this work, we test whether biologically inspired connectivity can improve image reconstruction in modern autoencoders by introducing a lightweight, biologically derived adapter that interfaces with a pre-trained VAE. The adapter modifies intermediate representations during reconstruction, improving image detail and fidelity without retraining the base model and with only a modest increase in parameters.
We evaluate our approach using the open-source Oasis Minecraft world generation model, which employs a modern Vision-Transformer-based VAE (ViT-VAE) to compress full-resolution images into a compact latent representation for efficient downstream processing. The VAE is trained on a large corpus of Minecraft images and achieves strong reconstruction fidelity within the distribution on which it was optimized.
As with most autoencoder-based compression systems, the VAE faces a fundamental tradeoff between efficiency and reconstruction quality. Compressing high-dimensional images into a constrained latent space inevitably discards information, often manifesting as over-smoothing, loss of fine texture, or reduced sharpness in reconstructed outputs. These effects become more pronounced when the model is evaluated on visually novel scenes or images that differ from the training distribution.
Common strategies to improve reconstruction quality include increasing model capacity, adjusting regularization, or retraining encoders and decoders. While effective, these approaches increase computational cost and limit deployability, particularly in systems where the VAE serves as an embedded component within a larger real-time pipeline. An alternative strategy is to augment the pretrained model with auxiliary, trainable components that act as lightweight “adapters.” Reservoir computing provides a theoretically appealing and computationally efficient framework for designing such adapters. Inspired by biological computation, reservoir computing leverages a fixed pool of nonlinear transformations, referred to as the reservoir, to project latent representations into a high-dimensional feature space where informative patterns can be more easily separated or amplified. In typical machine-learning workflows, the reservoir is randomly initialized and deliberately untrained, which makes training fast and stable because learning is confined to the readout.
To this end, we introduce a biologically derived adapter module that interfaces with intermediate VAE representations during reconstruction. The encoder and decoder weights remain frozen, and only the adapter parameters are trained. This design enables a controlled, parameter-matched comparison against typical (non-biologically derived) reservoir adapters and ensures that any observed improvements arise from the adapter itself rather than from increased model capacity or end-to-end optimization.
The structure and biological motivation of the adapter are described in the following section.
The adapter introduced in this work is derived from observed connectivity patterns in living neural networks cultured on multi-electrode arrays (MEAs) (Figure 1). These biological networks serve as a signal transformation substrate, producing structured spatiotemporal responses to controlled electrical stimulation. Rather than attempting to replicate biological computation in full, we extract specific, measurable properties of neural organization and translate them into reproducible computational primitives.

Figure 1. High-density multi-electrode array platform used for biological measurements. Cortical neurons are cultured within a well above a dense grid of 4,096 recording electrodes, forming active networks that span the electrode surface (right). The MEA supports precise electrical stimulation and simultaneous recording across the array, enabling measurement and modeling of spatial connectivity and spatiotemporal neural activity patterns used to construct the biologically derived adapters in this work.
Local connectivity is a defining feature of these networks. In our experiments, latent representations from the VAE are used to parameterize structured transformations informed by biological neural responses. We record the network’s response, defined as the spatiotemporal pattern of neuronal action potentials captured at sub-millisecond time scales across an array of 4,096 electrodes, and use these measurements to model how biological networks transform and propagate signals (Figure 2). These electrode-level models approximate how latent inputs would be processed under varying stimulation amplitude, spatial configuration, and inter-stimulation timing, revealing reliable patterns of local signal expansion and mixing despite substantial trial-to-trial variability (Figure 3).
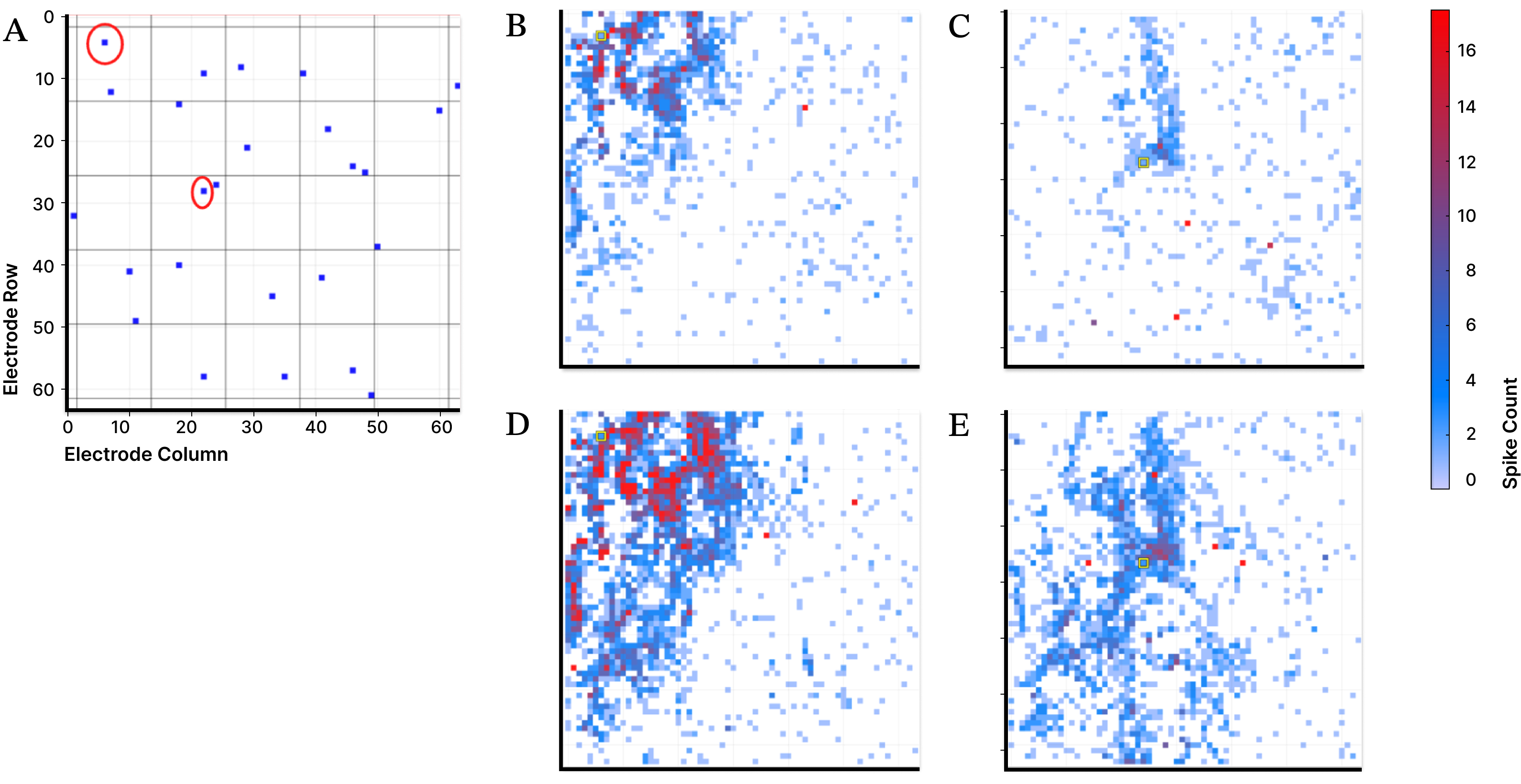
Figure 2. Electrical stimulation of individual electrodes in a MEA elicits localized neuronal responses that depend on stimulation site and input current. Cortical neurons are cultured on a 64 × 64 grid of surface electrodes, each of which can deliver current and record nearby action potentials. Panel A shows the locations of stimulated electrodes. Panels B and C show spatial patterns of evoked activity following moderate stimulation at two distinct sites, while panels D and E show responses at the same sites with higher input current. Increasing stimulation strength drives broader and higher-magnitude local activity, illustrating structured, site-dependent signal propagation across the network.
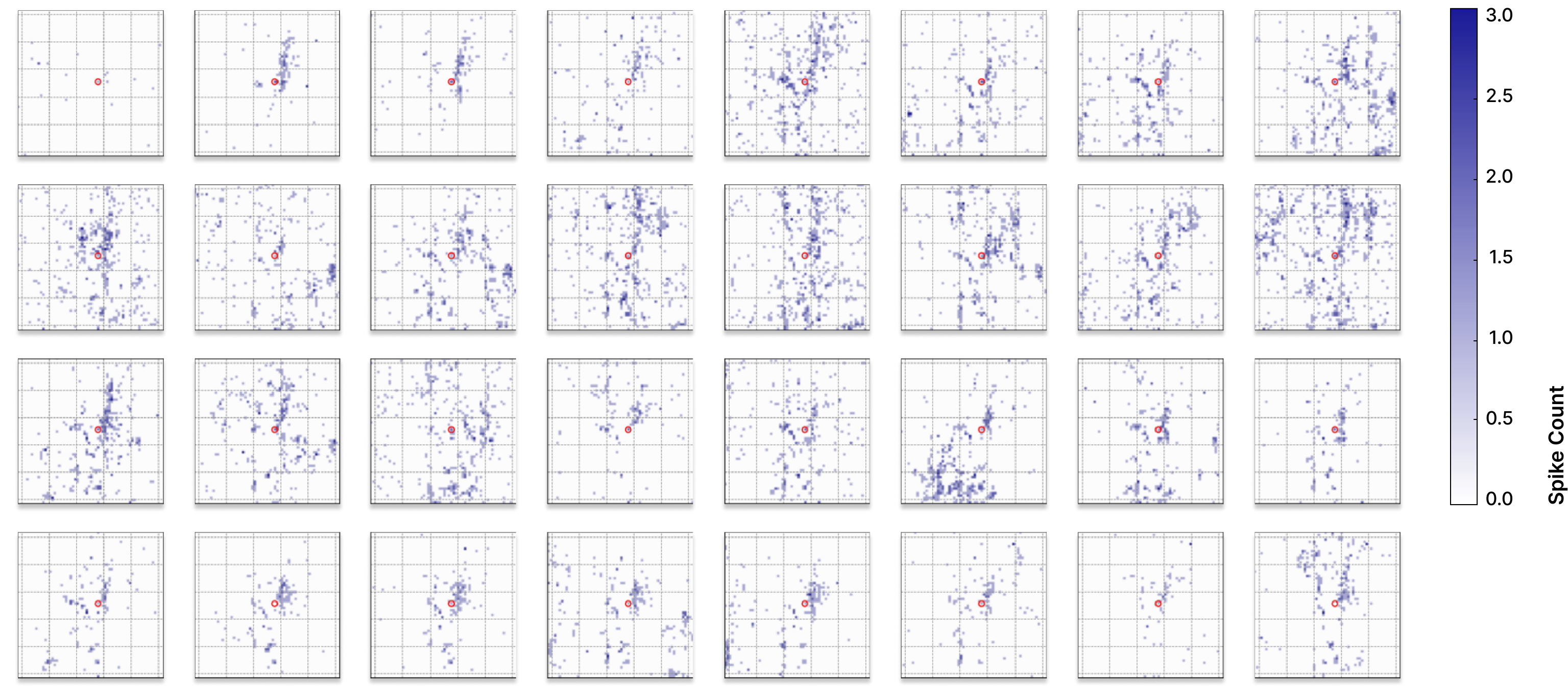
Figure 3. Trial-to-trial variability in biological neural responses. The same electrode is stimulated with the same input current across 32 trials, yet the resulting spatiotemporal activity patterns differ substantially. This intrinsic variability is a fundamental property of biological neural networks and highlights the challenge of integrating biological signals into machine-learning systems that typically assume consistent and precisely defined transformations.
To integrate these properties into a modern autoencoder, a biologically derived adapter that applies a structured transformation to intermediate VAE representations (Figure 4). The adapter is trained on 5,000 images drawn from five different video games, all resized to a common resolution. Latent representations from the frozen VAE encoder are passed through a neural filter informed by biologically measured or modeled connectivity patterns and implemented using standard PyTorch tooling. The transformed signal is mapped back into the native latent space through a lightweight, fully learnable projection layer and recombined with the existing VAE decoder.
During training, only the adapter parameters are optimized, while the base VAE remains unchanged. At deployment time, the system operates entirely in silico, with the biologically derived adapter functioning as a fixed transformation applied to intermediate representations. By introducing structured signal expansion informed by biological connectivity, the adapter improves image reconstruction quality without modifying the underlying VAE architecture or substantially increasing model capacity.
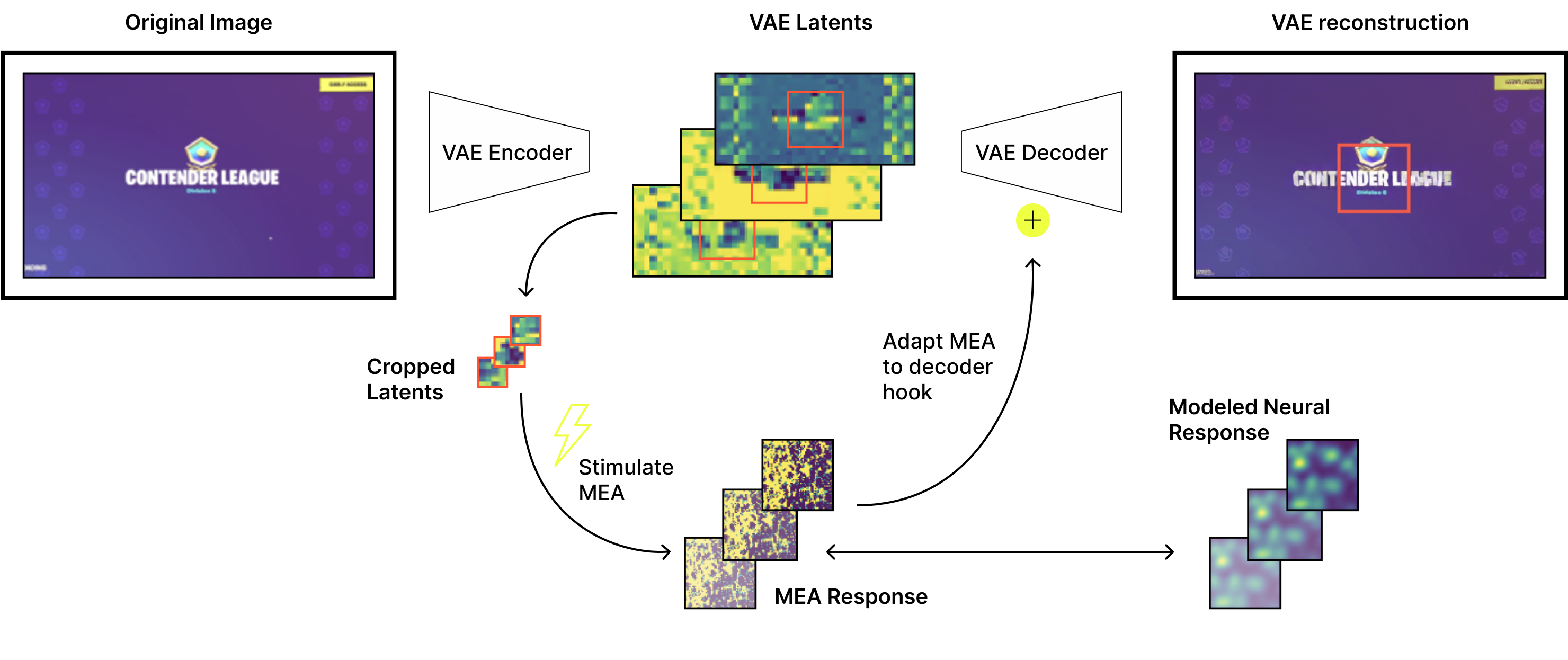
Figure 4. Overview of the biologically derived VAE adapter. An input image is compressed by the VAE encoder into a latent representation, and a selected subset of latent features is used to parameterize stimulation of an MEA. Observed or modeled MEA responses expand and redistribute latent information in a structured manner that reflects biological connectivity. This expanded signal is mapped back to the decoder-compatible latent space through a lightweight, learnable projection and reinserted into the VAE decoder. Experiments compare different MEA representations, stimulation parameters, and parameter-matched surrogate models to isolate the effect of the biologically informed transformation.
We evaluated the biologically derived adapter on image reconstruction using a VAE embedded within the Oasis pipeline. Reconstructions produced by the pre-trained VAE exhibit expected compression artifacts, including over-smoothing and loss of fine-grained structure, particularly in visually complex regions of the image. These artifacts reflect a common limitation of autoencoder-based compression when constrained latent representations must preserve high-frequency visual detail.
Integrating the biologically derived adapter into the reconstruction pathway consistently improves image quality (Figure 5). Qualitatively, reconstructed images retain sharper edges and clearer local structure compared to the baseline model. These improvements are achieved without modifying the VAE encoder or decoder and without retraining the base model, isolating the effect to the adapter itself.
Quantitatively, bio-inspired adapters outperform parameter-matched machine learning baselines. A first-order approximation of local biological connectivity achieves a 27% improvement in reconstruction quality relative to the baseline VAE, as measured by the relative decrease in reconstruction loss (Figure 5B). This improvement is obtained with an increase of 2.36 million parameters, or about 1% of the 229 million parameters in the base autoencoder. As a control experiment, we compared the biologically-derived adapter to a standard (non-biological) frozen reservoir with matched parameter counts, as well as a fully trainable reservoir that has additional degrees of freedom. When compared against parameter-matched control models, the biologically inspired adapter achieves loss values comparable to a fully trainable model.
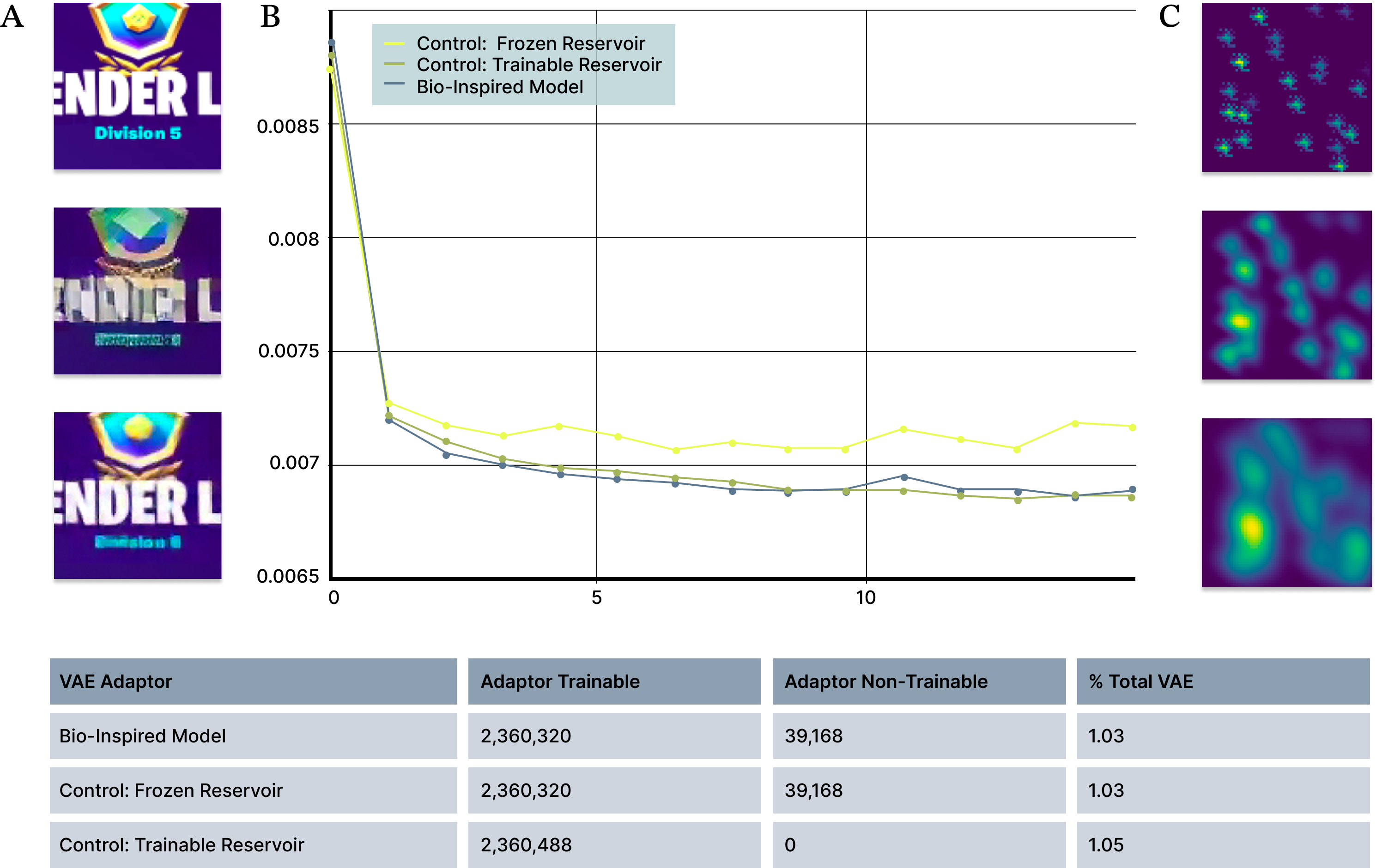
Figure 5. Bio-inspired adapters outperform matched-parameter machine-learning baselines. (A) Example image reconstructions showing improved perceptual quality with the bio-inspired adapter compared to the baseline model. Top: original image, Middle: reconstruction with baseline model; Bottom: reconstruction with bio-inspired adapter. (B) Reconstruction loss over training for bio-inspired, frozen-reservoir, and fully trainable reservoir models with matched parameter counts. The bio-inspired adapter consistently achieves lower loss than frozen reservoir models and approaches the performance of fully trainable models. (C) Example of a first-order approximation of local connectivity used in the bio-inspired adapter.
To identify which aspects of biologically inspired structure contribute most to reconstruction improvement, we systematically evaluated a range of spatial scales and response amplitudes in the bio-inspired models (Figure 5C). This controlled parameterization allowed us to distinguish connectivity patterns that improve decoding performance from those that degrade reconstruction quality.
Visual inspection of reconstructions suggests that the adapter helps preserve spatially localized information that is otherwise attenuated during compression. This behavior is consistent with the adapter’s design, which introduces structured signal expansion informed by biologically observed local connectivity patterns prior to encoding into the decoder-compatible latent space (Figure 4).
Finally, we extended this approach beyond bio-inspired models to adapters derived directly from measured biological activity. Using independently cultured neural networks on separate multi-electrode arrays, we generated MEA-based adapters by recombining previously recorded stimulation responses to match latent input patterns (Figure 6A–B). Despite substantial variability between the biological networks grown on each MEA, all MEA-derived adapters consistently outperform frozen reservoir baselines and approach the reconstruction quality achieved by fully trainable models (Figure 6C), indicating that biologically observed local connectivity provides a robust and transferable scaffold for improving representation quality under fixed architectural and compute constraints.
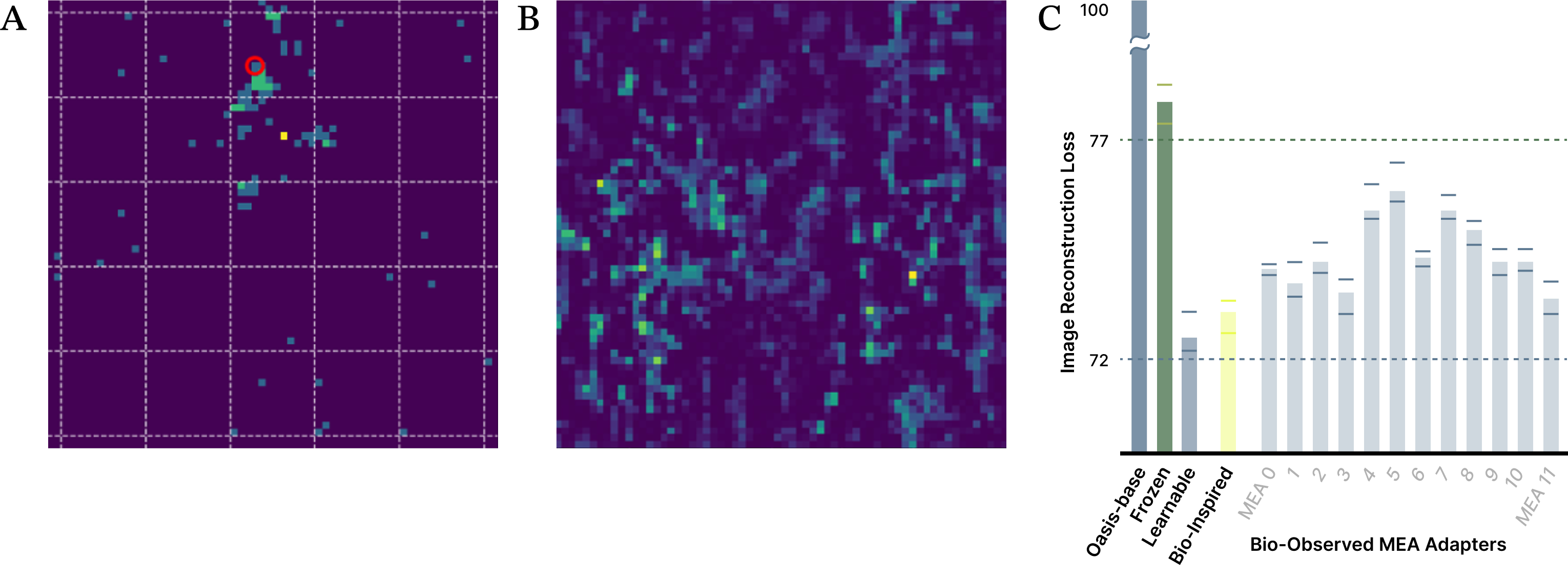
Figure 6. Adapters derived from measured biological responses outperform machine-learning frozen reservoir baselines. (A) Example neural response evoked by electrical stimulation of a single MEA electrode (red circle), with stimulation amplitude determined by the corresponding latent value from a local image region. Color indicates neural activity level. (B) Example MEA-wide representation of the center region of a training image, constructed by dynamically recombining previously measured stimulation responses according to latent location and amplitude. (C) Validation reconstruction loss for the bio-inspired model and 11 unique MEA-based adapters compared to matched-parameter frozen reservoir models, showing consistent improvement and performance approaching that of fully trainable models. Bars show mean loss across 8 random initializations, with min/max values for each condition shown as whiskers.
While these preliminary results suggest clear benefits from biologically derived adapters, several limitations remain. The current MEA-based approach operates within the spatial extent of a single array, and extending it to full-image adapters will require strategies for combining information across multiple MEAs and handling boundaries between arrays. Whether long-range biological connectivity across arrays is necessary, or whether local structure alone is sufficient, remains an open question.
There are also many ways to encode latent information into biological stimulation patterns. In this work, we use a simple mapping from latent amplitudes to stimulation strength and approximate full MEA responses by dynamically recombining previously measured stimulation events. While this enables generalization to new images, it likely omits nonlinear interactions between simultaneous stimulation sites, and understanding when such interactions matter will be important for future iterations.
More broadly, biological variability presents both challenges and opportunities. Neural responses vary across space and trials, and determining which aspects of this variability improve decoding performance versus those that degrade it remains an open problem. Exploring this design space, alongside choices in adapter architecture, stimulation channel selection, and down-stream artificial neural network hyperparameters, will be critical not only for improving reconstruction quality, but also for identifying general principles that transfer across models and tasks. These open questions motivate the progression of this blog series, where increasingly complex models are used to probe how biologically derived structure can inform learning dynamics and algorithm discovery.
We show that a lightweight, biologically derived adapter can improve image reconstruction quality in a variational autoencoder without retraining or architectural changes. By strengthening latent representations with minimal parameter overhead, this work provides a clear and measurable demonstration of how biologically informed structure can enhance core representation learning. Image reconstruction serves as a controlled test bed for algorithm discovery, and the improvements observed here directly informed the development of our adapter for diffusion-based video models, where the same principles translated to improved long-horizon stability. Within the broader arc of this series, these results illustrate how simple, interpretable settings can be used to systematically probe biological structure before extending those insights to more complex models and learning dynamics.
