 Explore our latest work
Explore our latest work

This blog series explores how biological neural networks can be integrated into modern AI systems. The posts progress from practical applications to increasingly general principles, moving from perception to representation, dynamics, and ultimately algorithm discovery. This first post focuses on perception, using computer vision as a controlled entry point for integrating biological neural dynamics into modern AI systems.
Note: This is a living document, and we are building in the open. We expect frequent updates and actively welcome feedback from the community.
For the first time, we show that living brain cells can be used to improve computer vision systems. By filtering images through biological neural networks before they reach a traditional AI model, the system learns to recognize images more efficiently than using silicon alone. This work shows how biological computing can begin with perception and scale toward more advanced AI capabilities.
When images were first processed by living brain cells and then passed to a standard AI model, the system classified images more accurately than when using AI alone. This improvement held even with a very simple AI model, showing that biological processing adds useful information rather than just extra complexity.

Biological neural activity enhances image classification. A handwritten digit (“5”) is transformed by a living neural network into a rich pattern of electrical activity (left), creating a representation with more features than the original image. When this neural representation is used to train a standard AI model, the model learns faster and reaches higher accuracy than when trained on the image alone, showing how biological dynamics make visual information easier for AI to use (right).
This work represents an early step in understanding how biological neural systems can complement today’s AI, starting with computer vision. We invite researchers, engineers, and collaborators interested in learning, perception, and the future of computing to engage with us as we build toward new approaches for how intelligent systems learn and adapt.
Computer vision has played a central role in the development of modern deep learning, serving as one of the earliest and most influential testbeds for neural network architectures. Benchmark datasets such as MNIST helped establish the feasibility of learning visual features from data, while more complex datasets like CIFAR pushed models toward handling greater variability and realism in visual inputs.
In this work, we revisit these canonical vision problems using a different computational substrate: living biological neural networks. Images are encoded into living neural systems using patterns of electrical stimulation, and the resulting neural activity is used as input to a downstream artificial neural network. We evaluate this approach on two datasets of increasing complexity, beginning with MNIST, which consists of simple handwritten digits that can be directly encoded, and then extending to CIFAR, which contains natural images that require a learned feature encoding prior to biological stimulation.
By comparing performance across these benchmarks, we assess how biological neural dynamics reshape visual representations and whether this transformation provides practical benefits for image classification across different levels of visual complexity.
We use living biological neural networks grown on multielectrode arrays as a computational substrate for processing visual information (Figure 1). These networks self-organize into recurrent, distributed systems that process inputs through analogue electrical activity, offering a mode of computation that differs fundamentally from digital, feedforward operations.
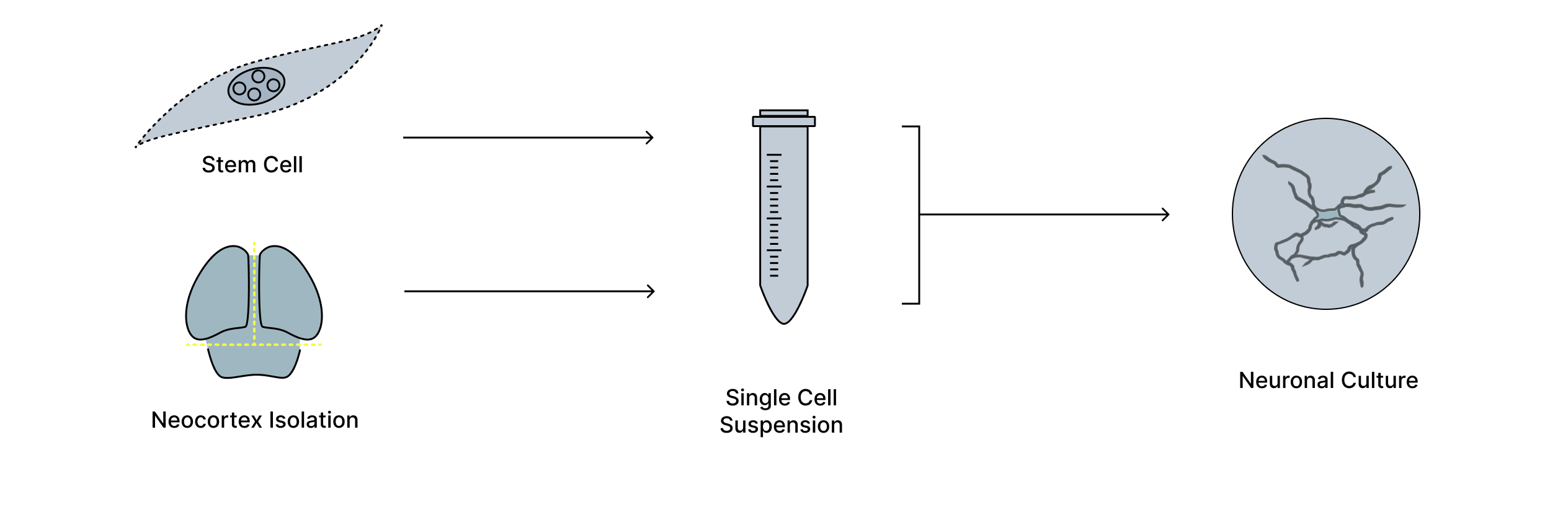

Figure 1. Generation of living neuronal cultures on electrode arrays. Neurons are derived from biological tissue, isolated, and dissociated into a single-cell suspension before being grown on high-density multielectrode arrays. Over time, these cells form living neuronal cultures that can be interfaced with electronic systems, providing a physical substrate for studying and harnessing neural activity.
Visual inputs are translated into patterns of electrical stimulation delivered across the electrode array. Once stimulated, the biological network transforms these inputs through its intrinsic neural dynamics, producing spatiotemporal patterns of activity that reflect interactions both within and beyond the directly stimulated region.
We record the resulting neural activity and summarize it using simple measures of spike rate over short time windows. This intentionally minimal readout is then used as input to a conventional artificial neural network for image classification, allowing us to isolate the contribution of biological neural dynamics without relying on complex decoding strategies (Figure 2).
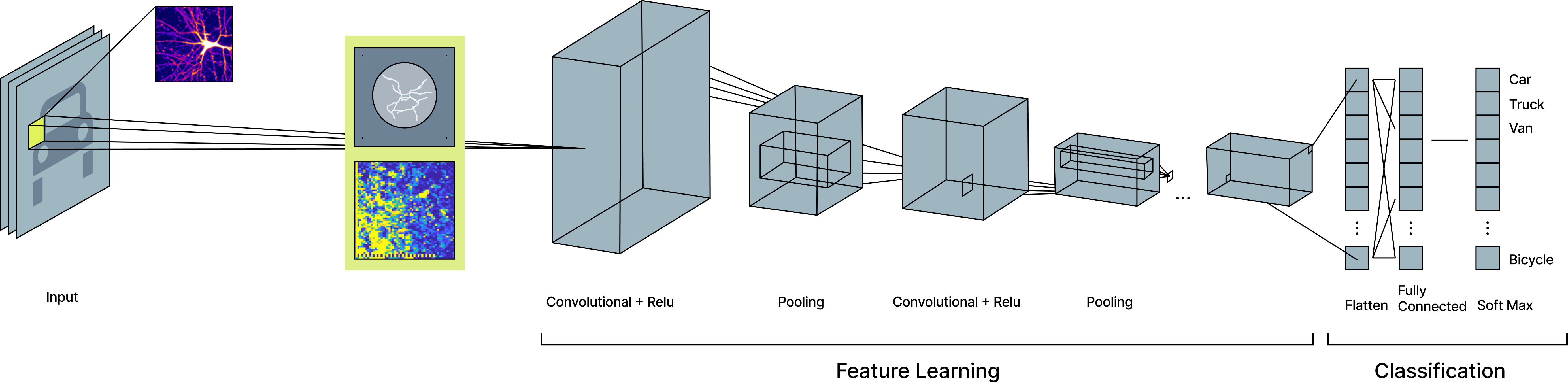
Figure 2. Biological preprocessing of MNIST images for classification. MNIST images are first encoded as electrical stimulation patterns and passed through a living neural network, which transforms the image into a richer neural representation. This biological representation is then provided as input to a simple convolutional neural network, where standard feature learning and classification are performed. By inserting biological processing before conventional feature extraction, the downstream model learns from a representation that contains more complex structure than the original image alone.
To establish a clear baseline for how biological neural dynamics process visual information, we first applied our approach to the MNIST dataset, which consists of simple grayscale images of handwritten digits well suited for direct biological encoding.
We selected a limited subset of the MNIST dataset to avoid saturating the downstream classifier and binarized the MNIST digits by converting grayscale pixels into white digits on a black background. These binarized images were spatially centered on a 64 × 64 multielectrode array populated with a living neural network, and electrical stimulation was delivered to electrodes corresponding to the white pixels of each digit.
Following stimulation, the biological neural network responded with rich, nonlinear spatiotemporal patterns of activity across the electrode array. We recorded this activity over a short time window and summarized it using spike-rate measurements, yielding a neural representation of each digit that extended beyond the directly stimulated region. These neural responses were then used as input to a downstream artificial neural network for classification (Figure 3, left).
To isolate the contribution of biological processing, the downstream silicon network was intentionally kept simple. It consisted of an input layer receiving either the binarized MNIST images or the corresponding neural responses, followed by a single convolutional layer with five 4 × 4 kernels, a linear layer followed by a ReLU nonlinearity, then a classification layer for the ten MNIST digit classes. Both inputs were trained under identical conditions, allowing a direct comparison of classification performance with and without biological preprocessing.
We first evaluated whether biological neural preprocessing improves image classification on the MNIST dataset. Training a convolutional neural network on biological neural responses led to both faster learning and higher final classification accuracy compared to training on binarized MNIST images alone (Figure 4). Because the downstream model architecture and number of trainable parameters were identical with or without neural pre-processing these improvements reflect differences in the input representation rather than increased model capacity.
To understand how biological processing altered the representation, we compared binarized MNIST digits, their corresponding neural responses, and downstream classifier activations (Figure 3, right). When MNIST digits were embedded into the biological network, neural activity extended beyond electrodes directly stimulated by the digit, indicating that recurrent biological dynamics distributed information across the network.
Analysis of convolutional layer activations revealed that learning relied on features distributed well outside the original digit location (Figure 3). Visualizing the most extreme activation changes highlighted consistent, class-specific patterns beyond the stimulated region, suggesting that biological neural dynamics expanded the effective feature space available to the classifier.
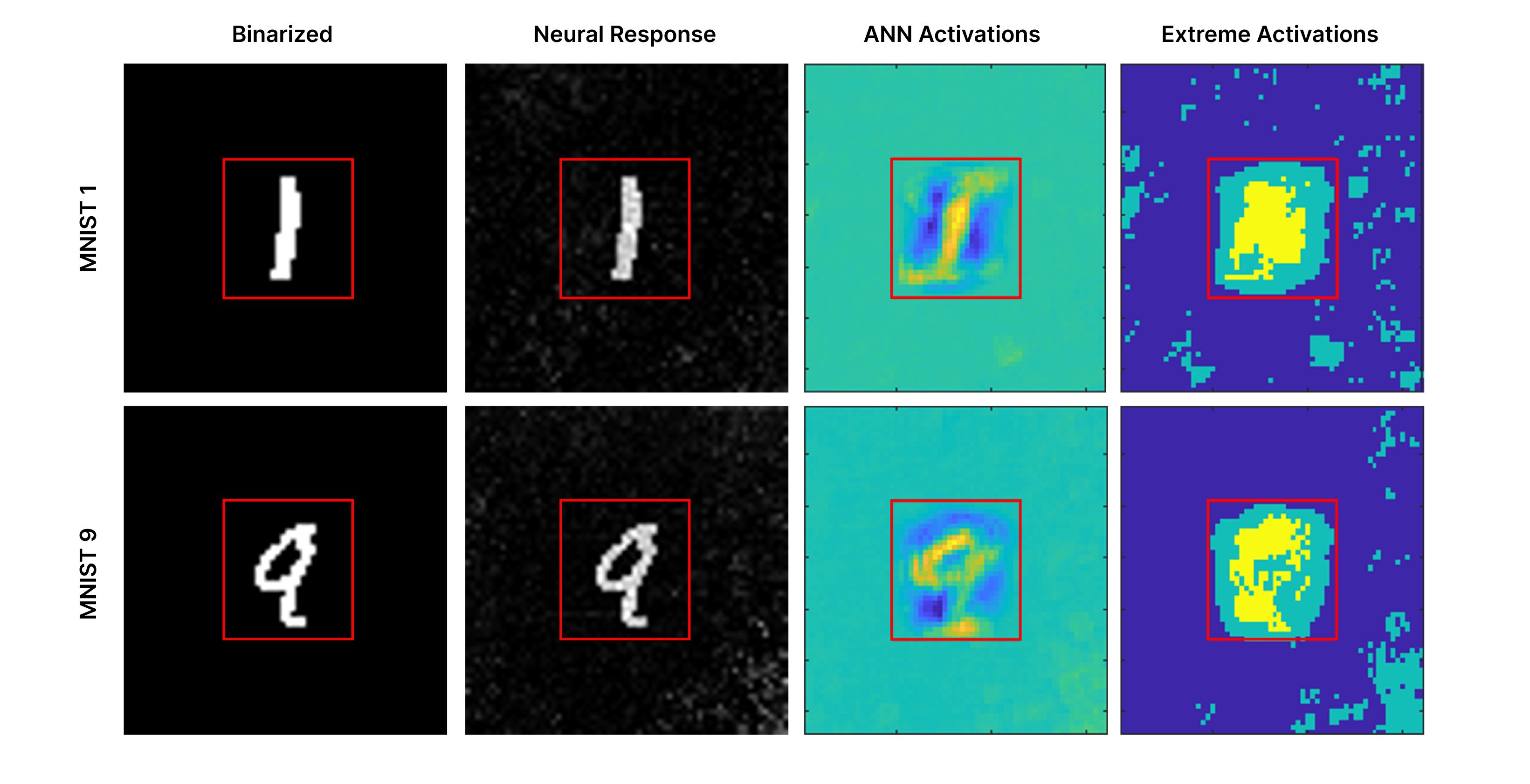
Figure 3. Biological neural dynamics project images into a higher dimensional space leveraged for classification. MNIST digits are binarized and used to stimulate a biological neural network (left), producing neural responses that extend beyond the region of direct stimulation (second column; red box indicates the original 28×28 digit area). When a convolutional neural network is trained on binarized MNIST images alone, the resulting changes in convolutional layer activations are shown in the third column (ANN activations), reflecting features learned from biological inputs. The most extreme activations from that network are shown in the fourth column, with the top 5% of changes highlighted in yellow and the top 20% in aqua. These biologically driven activation patterns reveal consistent, class-specific structure outside the original stimulation region that the classifier leverages to improve performance.
To further probe the source of this information, we trained classifiers on different subsets of the neural signal. As expected, pre-stimulation activity performed at chance. In contrast, post-stimulation neural responses enabled accurate classification when using signals from the full electrode array, the directly stimulated region alone, or even regions outside the stimulation area (Figure 4, right).
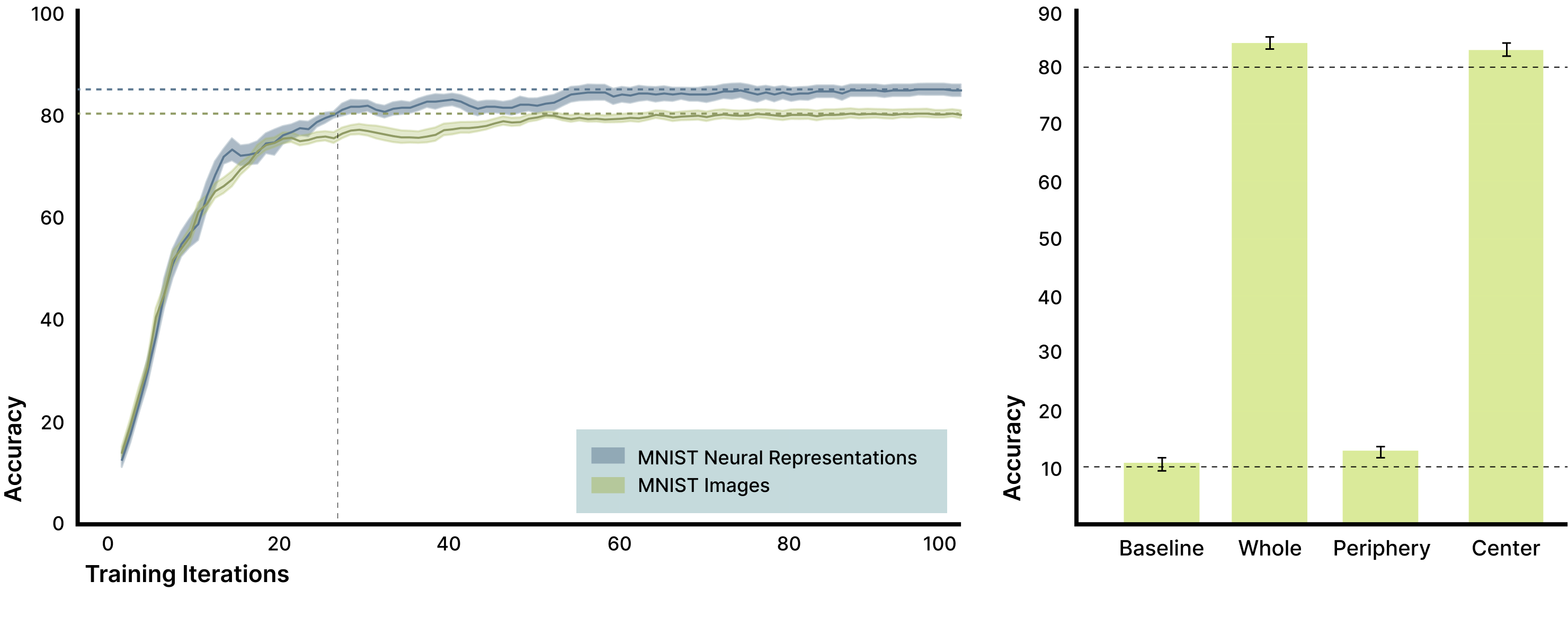
Figure 4. Biological neural responses improve MNIST classification and reveal distributed signal structure. A convolutional neural network trained on biological neural responses achieved higher classification accuracy than the same network trained on binarized MNIST images alone, with an average improvement of 4.7% after ten training epochs (left; shaded regions show SEM across ten network initializations). Training classifiers on different components of the neural response (right) shows that activity from the full electrode array (“Whole”) and the directly stimulated region (“Center”) both support high classification accuracy, while activity from regions outside the stimulation area (“Periphery”) performs reliably above chance (approximately 13% versus 10% chance). This confirms mechanistically that biologically induced, distributed features visible as activation changes outside the original image footprint contribute meaningful information that the classifier leverages to improve performance.
Together, these results show that biological neural networks transform simple visual inputs into higher-dimensional, distributed representations that make downstream learning easier, rather than merely increasing computational complexity.
To test whether the benefits observed for MNIST extend to more complex visual inputs and to assess the scalability of the approach, we next applied our pipeline to the CIFAR dataset. Unlike MNIST, CIFAR images contain natural scenes with greater variability in texture, object identity, viewpoint, and background, making direct pixel-level encoding insufficient and requiring an additional feature extraction step before biological stimulation.
To address this, we first trained a convolutional neural network on the full CIFAR training set to learn feature representations useful for classification. After training, the learned convolutional filters were fixed and used as feature extractors, transforming each image into a small set of low-dimensional activation patterns. These activations were binarized and mapped onto the multielectrode array, yielding four spatial stimulation patterns per image.
Each CIFAR image was represented by a sequence of these stimulation patterns delivered to the biological network. Following each stimulation, neural activity was recorded over a short post-stimulation window and summarized using spike-rate measurements, producing a set of biological neural responses corresponding to the original image.
These neural responses were concatenated and used as input to a downstream convolutional classifier, which was trained either on the biological neural responses or directly on the stimulation patterns alone for comparison. By holding the learned visual features and downstream model architecture fixed and varying only whether biological processing was applied, this encoding strategy isolates the contribution of biological neural dynamics when processing complex natural images.

Figure 5. Encoding CIFAR images into stimulation patterns for biological networks. After training a convolutional neural network on the CIFAR dataset, the learned filters were fixed and used as feature extractors to process CIFAR images. Each image was transformed into a small set of class-relevant feature maps that were binarized using a thresholding step, producing four binary stimulation patterns per image. These binary patterns defined the corresponding stimulation electrodes on the multielectrode array.
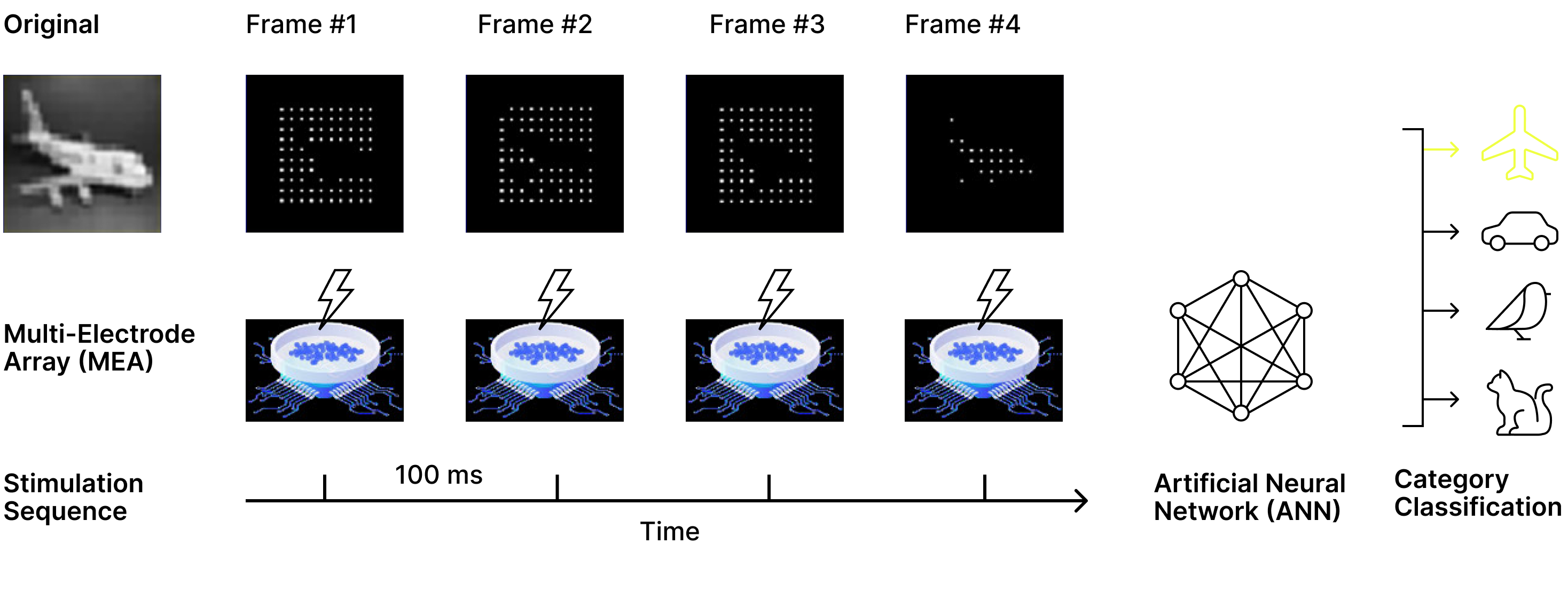
Figure 6. Sequential stimulation and biological readout for CIFAR classification. Each CIFAR image was represented by four binary stimulation patterns that were delivered sequentially to the biological neural network. Following each stimulation, neural activity was recorded to produce four biological response patterns per image. These neural responses were concatenated and used as input to a downstream convolutional classifier, which was trained either on the biological neural signals or directly on the stimulation patterns alone to isolate the contribution of biological neural dynamics.
We evaluated the effect of biological neural processing on CIFAR images using the encoding strategy described above. A central result of this experiment is that complex, natural images can be successfully encoded into structured stimulation patterns and interfaced with a biological neural network using a multielectrode array. Despite the increased variability and visual complexity of CIFAR images compared to MNIST, this approach produced stable and repeatable neural responses suitable for downstream classification.
When a convolutional classifier was trained on biological neural responses, performance reached the same final classification accuracy as a classifier trained directly on the corresponding stimulation patterns, while achieving higher accuracy earlier in training (Figure 7). However, models trained on biological neural responses converged approximately three times faster, reaching peak performance in substantially fewer training iterations. Because the downstream model architecture and visual features were identical in both cases, this acceleration reflects the contribution of biological neural dynamics rather than differences in model capacity, and implies a threefold reduction in the computational and energy demands required to train the classifier to peak performance.
Importantly, these results also demonstrate flexibility in the order of operations between biological and artificial neural systems. In the CIFAR setting, an artificial neural network was first used to extract structured visual features, which were then processed by a biological network before final classification. This contrasts with the MNIST experiments, where biological processing operated directly on raw visual inputs.
Together, the CIFAR results show that biological neural networks can be integrated at different stages of the vision pipeline depending on task complexity and design constraints. This composability suggests that biological processing can serve as a flexible computational transform, motivating its application beyond perception and toward more general forms of representation learning and dynamical modeling.
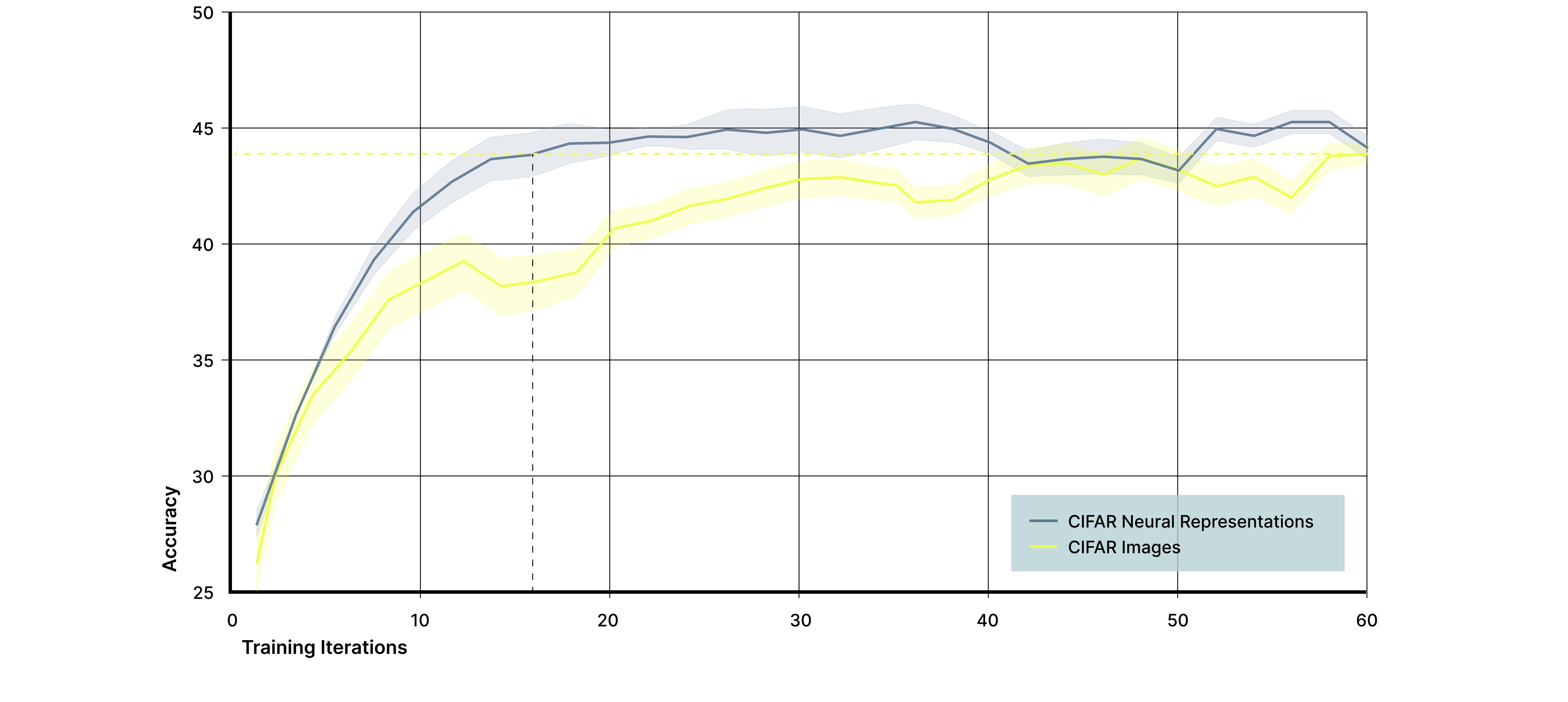
Figure 7. Biological neural responses accelerate learning on CIFAR images. A convolutional neural network was trained to classify four CIFAR object categories using either biological neural responses (blue) or the corresponding stimulation patterns alone (yellow) as input. While both input signals reached a similar final classification accuracy of approximately 43.5%, models trained on biological neural responses converged roughly three times faster, demonstrating that biological neural dynamics reshape complex visual inputs in a way that accelerates downstream learning.
This work represents an initial step in integrating biological neural networks into computer vision pipelines, with several limitations remaining. For MNIST, our experiments were intentionally small in scale and designed to establish mechanism rather than optimize performance. Future work will evaluate larger datasets and additional neural cultures to assess robustness, variability, and scaling behavior, particularly to better understand how neural activity outside the stimulated region contributes to classification.
For CIFAR, we demonstrate that complex natural images can be encoded into multielectrode arrays and that biological processing accelerates learning. Further improvements are likely through alternative encoding strategies, including transformer-based vision models, to better capture structure in higher-resolution and more diverse images.
Across both datasets, our results rely on simple spike-rate summaries of neural activity, which do not capture the temporal structure known to be central to biological computation. Ongoing work focuses on decoding spike timing and using time as an explicit encoding dimension, leveraging the ability of biological networks to encode and integrate information across sequential inputs. More broadly, these results show that the order of operations between biological and artificial neural systems is flexible, enabling biological processing to operate at different stages of the pipeline depending on task demands. This flexibility provides both a practical path for integration and a controlled framework for probing how biological dynamics transform representations.
We show that biological neural networks can be integrated into computer vision pipelines to reshape visual representations in ways that improve downstream learning, from simple handwritten digits to complex natural images. By demonstrating that visual information can be encoded into multielectrode arrays and processed by biological networks, and that biological and artificial systems can be flexibly composed in different orders, this work establishes perception as a practical entry point for biological computing. Building on this foundation, subsequent work extends these ideas beyond vision toward representation learning, temporal generative models, and ultimately the use of real neural dynamics as a platform for algorithm discovery.
